Nuclear Reactions Rates and Core Boundary Mixing on the Seismology of Red Clump Stars
For the complete introduction to this lab and to red clump stars, go to the beginning of Lab 2.
Maxilab2: Exploring Nuclear Reaction Rates and Networks in MESA
Background:
During the core helium-burning (CHeB) phase, the energy production from the core is primarily driven by two nuclear reactions:
- The triple-alpha process ( ): three helium nuclei ( ) fuse to form carbon ( ). It occurs in two steps: and .
- The reaction: a carbon nucleus captures an alpha particle to form oxygen ( ).
As mentioned in the introduction to the Maxilab, nuclear reaction rates and their temperature dependence are a source of uncertainty in RC evolution, leading to different stellar interior structures and different period spacings to compare with observations. Helium burning in RC stars is driven primarily by two nuclear reactions. The process dominates during the early stages of the core helium-burning phase, while the reaction becomes more important later on.
These reactions are highly temperature-sensitive and uncertain, especially the rate. In stellar interiors, this reaction occurs at low energies (hundreds of keV), where the cross-section is extremely small because the Coulomb barrier suppresses the reaction probability. Direct measurements in the laboratory at these energies are impractical, so experiments are performed at higher energies (in the MeV range) and extrapolated to stellar conditions using theoretical models. This extrapolation introduces significant uncertainty.
Changes in these rates affect the core composition, the duration of the core helium-burning (CHeB) phase, and, indirectly, the size and structure of the convective core. This, in turn, influences the period spacing , making reaction rates a key variable to explore.
In this Maxilab2, we will learn how to change the reaction rates in MESA, and we will recover part of Figure 5 and Figure 6 from Noll et al. 2024 together as a class.
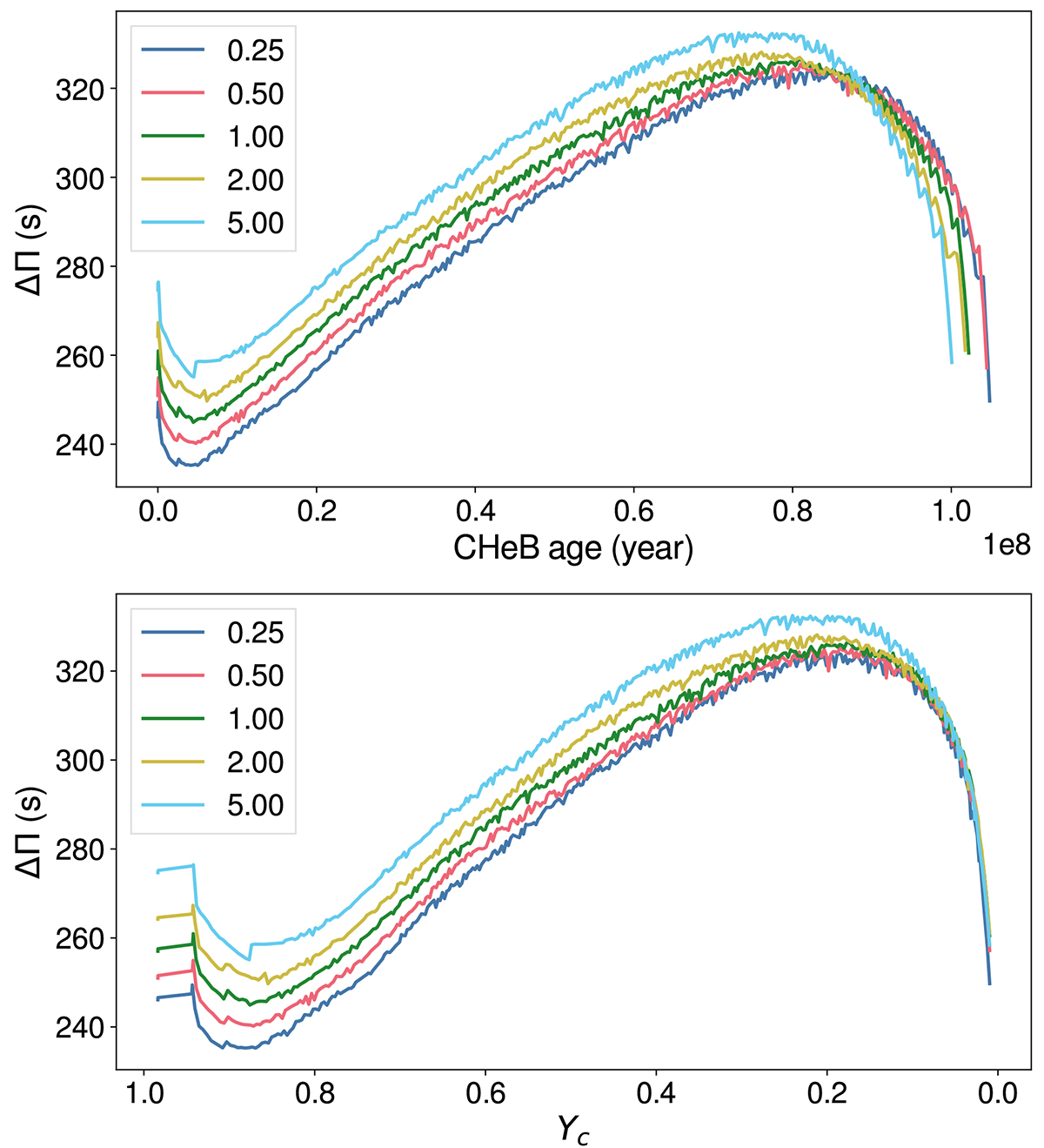
Figure 5 from Noll et al. 2024 shows the evolution of
during the CHeB phase for different
reaction rates as a function of CHeB age (top panel) and central helium abundance (lower panel).
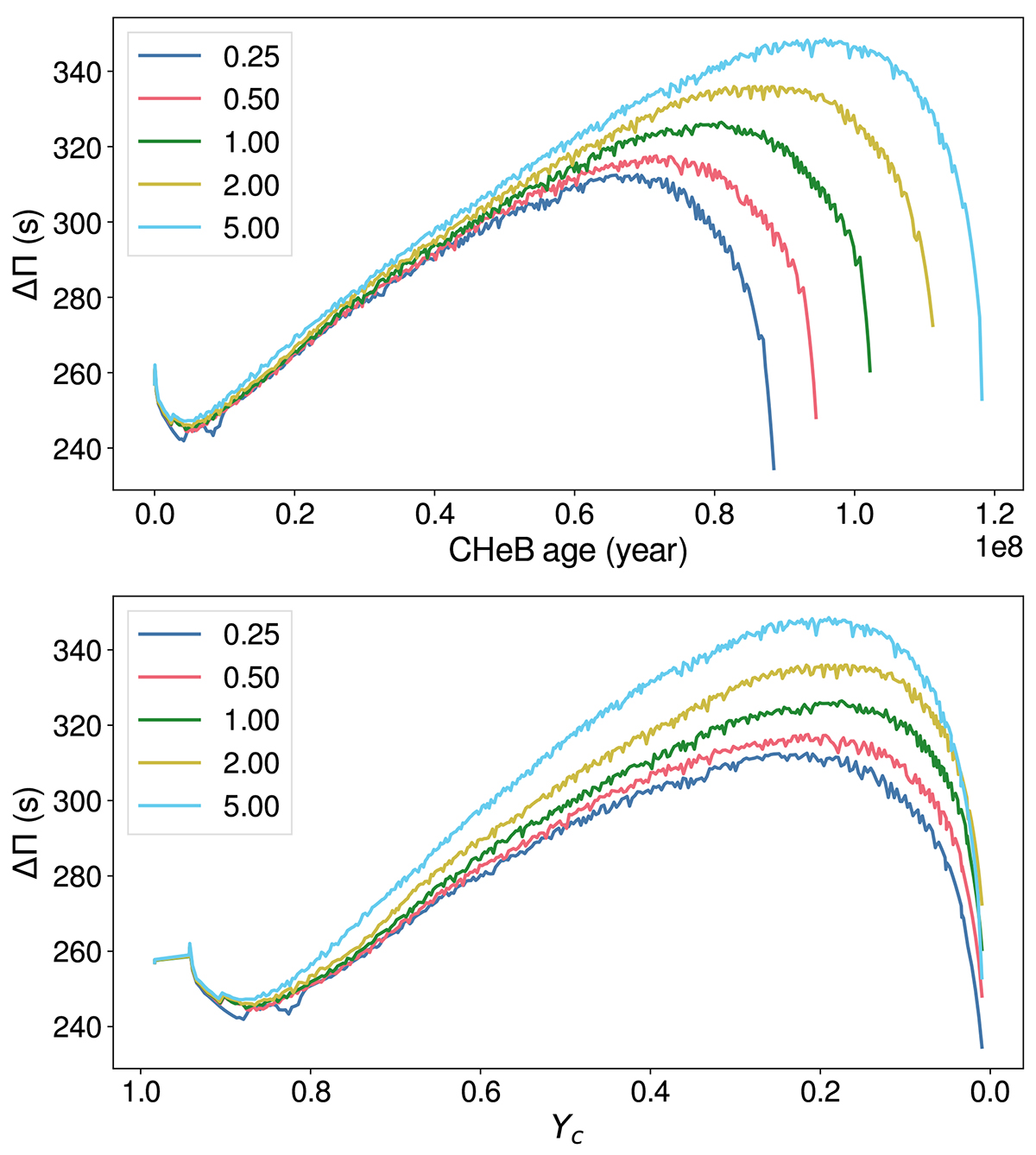
Same as Figure 5, but for different
reaction rates:
Aims
MESA aims: : In this Maxilab, you will learn how to find and modify the nuclear reaction network, how to read the reaction network files, and where to find them in $MESA_DIR. You will learn how to change reaction rates for specific profiles and how to incorporate all of that into your MESA inlist. You will also learn how to print global MESA parameters in the terminal by making a small edit to the run_star_extra file. Finally, you will learn how to navigate the MESA documentation.
Science aims: You will learn how changing the reaction rate for a specific evolutionary phase in stellar evolution affects the composition and inner structure, and hence the period spacing pattern.
Solution: In case you get stuck at any point during the exercises, you can find the solution for this Maxilab here.
Setting up the MESA model
In this Maxilab, we will fix the mixing scheme to maximal overshooting to test the effect of reaction rates on the period spacing.
If you ran your MESA models in Maxilab1 using the maximal overshooting scheme, you can copy that work directory and continue this Maxilab with it. If you used another mixing scheme, or if you are not sure about your solution, please download the starting work directory from here. At this link, you can download a zipped directory named maxilab2.zip.
If you want to unzip the folder using the terminal, you can use:
unzip maxilab2.zipTerminal commands:
cd maxilab2
./clean && ./mkMake sure that you are able to compile and start the run without any issues. Stop the run after you see the pgplot window.
Task 1: Identify and Inspect the Default Nuclear Reaction Network
- Look for the name of the nuclear reaction network that MESA uses by default.
- Locate the corresponding network file, open it, and find the reaction rate entries related to the He-burning processes.
Hint 1
- Look for the
&star_jobcontrols in the MESA documentation (available here, underReference and Defaults -> star_job). Find the default nuclear reaction network name there. - Check your
inlistfor the&star_jobcontrols that set the reaction network. If none is specified, MESA will use its default. - Look in
$MESA_DIR/data/net_data/nets/to see which file corresponds to that default network. - Open the network file in a text editor and examine the listed reaction entries.
- Identify the entries corresponding to the triple-alpha process and the reaction.
- Copy the names of these entries for
Task 2.
Solution 1
- The name of the default nuclear reaction network in MESA is
basic.net. This is also the file you need to open to find the entries corresponding to the triple-alpha process and the reaction, located in$MESA_DIR/data/net_data/nets/. - The entries are:
r_he4_he4_he4_to_c12,r_c12_ag_o16.
The names of the reaction rate entries you found refer to files that describe the reaction rate of each process as a function of temperature. If you want to use another file for a specific process, for example one computed in a recent study that MESA does not include yet, you can add these files and direct MESA to read them. These files can be found under: ls $MESA_DIR/data/rates_data/rate_tables. We will not change these files directly during this lab, but will change the reaction rates directly through the inlist.
Task 2: Assign and Test Special Reaction Rate Factors
- Find the
inlistvariables that need to be changed in order to modify the reaction rates for the two He-burning processes (use the names of the reaction rate entries you found inTask 1) and edit yourinlistaccordingly. - The reaction rate values to test are the default rates multiplied by x - (for example, 0.25, 0.5, 1, 2, and 5) for the process while keeping the reaction rate at 1, and vice versa. Each table will be assigned a different value to compute.
To pick a value, go to this Google Sheet (local copy) and write your name on one of the rows, where each row has a value for the and reaction rates. Update your inlist accordingly. Later in this lab, we will print the maximum value of and the stellar age at which it occurs, and we will plot it live for the different values in the table. After you finish your run, you can update the value in the sheet.
Hint 2
- Check the
&star_jobsection in the MESA documentation for variables that let you specify special reaction rate factors for individual reactions. You can also look in the$MESA_DIR/star/defaults/star_job.defaultsfile. - Look for three
inlistvariables that let you control the number of special rate factors to assign, specify the name of the reaction, and set the scaling factor.
Solution 2
This is how the addition to the &star_job section in your inlist_project file should look:
num_special_rate_factors = 2
reaction_for_special_factor(1) = 'r_he4_he4_he4_to_c12'
reaction_for_special_factor(2) = 'r_c12_ag_o16'
special_rate_factor(1) = 1 ! 0.25, 0.5, 1.00, 2.00, 5.00
special_rate_factor(2) = 5 ! 0.25, 0.5, 1.00, 2.00, 5.00In this example, the
reaction rate (r_he4_he4_he4_to_c12) is set to 1, and the
reaction rate (r_c12_ag_o16) is scaled by a factor of 5.
Task 3: Enable Reaction Network Printing in Terminal
Find the two &star_job variables that will print to the terminal during the run: one that lists the reactions in the current network, and one that provides detailed information about those reactions.
Solution 3
This is how the addition to the &star_job section in your inlist_project file should look:
show_net_reactions_info = .true.
list_net_reactions = .true.Task 4: Tracking the Peak Value of delta_Pg during Evolution
Your task is to modify run_star_extras.f90 so that during the run:
- It finds the maximum value of
delta_Pgever reached during the evolution. - Each time a new peak is found, it prints the value of delta_Pg to the terminal along with the stellar age at that step.
- By the end, the last printed value will be the global maximum.
We want to set a parameter for the global peak of delta_Pg, which we will call peak_delta_Pg. We need to declare it so that during the run, peak_delta_Pg is not overwritten at each time step in MESA unless the new value of delta_Pg is higher than the previous one. We will declare this parameter as a real number with double precision.
Hint 4.1
delta_Pg and the stellar age values at each step by using the star_info object: s% delta_Pg and s% star_age.Hint 4.2
You need to modify run_star_extras.f90 in three places:
- We will declare
peak_delta_Pgat the top of ourrun_star_extras.f90. - We will modify
extras_startuproutine — runs once at the very start. - We will modify
extras_finish_step— runs after every timestep.
Solution 4
- We will declare
peak_delta_Pgat the beginning ofrun_star_extra.f90file. Addreal(dp) :: peak_delta_Pgafterimplicit none. - Look for the
extras_startuproutine and initializepeak_delta_Pgparameter:
subroutine extras_startup(id, restart, ierr)
integer, intent(in) :: id
logical, intent(in) :: restart
integer, intent(out) :: ierr
type (star_info), pointer :: s
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
! initialize peak_delta_Pg
write(*, *) 'initializing peak_delta_Pg'
peak_delta_Pg = 0
end subroutine extras_startup- Look for the
extras_finish_stepand add the following if condition:
integer function extras_finish_step(id)
integer, intent(in) :: id
integer :: ierr
type (star_info), pointer :: s
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
extras_finish_step = keep_going
! to save a profile,
! s% need_to_save_profiles_now = .true.
! to update the star log,
! s% need_to_update_history_now = .true.
!! Task block begins here:
!write(*,*) 'delta_Pg:', s% delta_Pg
if (s% delta_Pg > peak_delta_Pg) then
peak_delta_Pg = s% delta_Pg
write(*,*) 'New peak of delta_Pg found:', peak_delta_Pg
write(*,*) 'star age:', s%star_age
end if
!! TASK block ends here
! see extras_check_model for information about custom termination codes
! by default, indicate where (in the code) MESA terminated
if (extras_finish_step == terminate) s% termination_code = t_extras_finish_step
end function extras_finish_stepIf you want to see the complete run_star_extra.f90 file, click here.
! ***********************************************************************
!
! Copyright (C) 2010-2019 Bill Paxton & The MESA Team
!
! this file is part of mesa.
!
! mesa is free software; you can redistribute it and/or modify
! it under the terms of the gnu general library public license as published
! by the free software foundation; either version 2 of the license, or
! (at your option) any later version.
!
! mesa is distributed in the hope that it will be useful,
! but without any warranty; without even the implied warranty of
! merchantability or fitness for a particular purpose. see the
! gnu library general public license for more details.
!
! you should have received a copy of the gnu library general public license
! along with this software; if not, write to the free software
! foundation, inc., 59 temple place, suite 330, boston, ma 02111-1307 usa
!
! ***********************************************************************
module run_star_extras
use star_lib
use star_def
use const_def
use math_lib
use num_lib, only: find0, two_piece_linear_coeffs
implicit none
real(dp) :: peak_delta_Pg
! these routines are called by the standard run_star check_model
contains
subroutine extras_controls(id, ierr)
integer, intent(in) :: id
integer, intent(out) :: ierr
type (star_info), pointer :: s
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
! this is the place to set any procedure pointers you want to change
! e.g., other_wind, other_mixing, other_energy (see star_data.inc)
! the extras functions in this file will not be called
! unless you set their function pointers as done below.
! otherwise we use a null_ version which does nothing (except warn).
s% extras_startup => extras_startup
s% extras_start_step => extras_start_step
s% extras_check_model => extras_check_model
s% extras_finish_step => extras_finish_step
s% extras_after_evolve => extras_after_evolve
s% how_many_extra_history_columns => how_many_extra_history_columns
s% data_for_extra_history_columns => data_for_extra_history_columns
s% how_many_extra_profile_columns => how_many_extra_profile_columns
s% data_for_extra_profile_columns => data_for_extra_profile_columns
s% how_many_extra_history_header_items => how_many_extra_history_header_items
s% data_for_extra_history_header_items => data_for_extra_history_header_items
s% how_many_extra_profile_header_items => how_many_extra_profile_header_items
s% data_for_extra_profile_header_items => data_for_extra_profile_header_items
end subroutine extras_controls
subroutine extras_startup(id, restart, ierr)
integer, intent(in) :: id
logical, intent(in) :: restart
integer, intent(out) :: ierr
type (star_info), pointer :: s
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
! initialize peak_delta_Pg
write(*, *) 'initializing peak_delta_Pg'
peak_delta_Pg = 0
end subroutine extras_startup
integer function extras_start_step(id)
integer, intent(in) :: id
integer :: ierr
type (star_info), pointer :: s
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
extras_start_step = 0
end function extras_start_step
! returns either keep_going, retry, or terminate.
integer function extras_check_model(id)
integer, intent(in) :: id
integer :: ierr
type (star_info), pointer :: s
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
extras_check_model = keep_going
if (.false. .and. s% star_mass_h1 < 0.35d0) then
! stop when star hydrogen mass drops to specified level
extras_check_model = terminate
write(*, *) 'have reached desired hydrogen mass'
return
end if
! if you want to check multiple conditions, it can be useful
! to set a different termination code depending on which
! condition was triggered. MESA provides 9 customizable
! termination codes, named t_xtra1 .. t_xtra9. You can
! customize the messages that will be printed upon exit by
! setting the corresponding termination_code_str value.
! termination_code_str(t_xtra1) = 'my termination condition'
! by default, indicate where (in the code) MESA terminated
if (extras_check_model == terminate) s% termination_code = t_extras_check_model
end function extras_check_model
integer function how_many_extra_history_columns(id)
integer, intent(in) :: id
integer :: ierr
type (star_info), pointer :: s
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
how_many_extra_history_columns = 0
end function how_many_extra_history_columns
subroutine data_for_extra_history_columns(id, n, names, vals, ierr)
integer, intent(in) :: id, n
character (len=maxlen_history_column_name) :: names(n)
real(dp) :: vals(n)
integer, intent(out) :: ierr
type (star_info), pointer :: s
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
! note: do NOT add the extras names to history_columns.list
! the history_columns.list is only for the built-in history column options.
! it must not include the new column names you are adding here.
end subroutine data_for_extra_history_columns
integer function how_many_extra_profile_columns(id)
integer, intent(in) :: id
integer :: ierr
type (star_info), pointer :: s
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
how_many_extra_profile_columns = 0
end function how_many_extra_profile_columns
subroutine data_for_extra_profile_columns(id, n, nz, names, vals, ierr)
integer, intent(in) :: id, n, nz
character (len=maxlen_profile_column_name) :: names(n)
real(dp) :: vals(nz,n)
integer, intent(out) :: ierr
type (star_info), pointer :: s
integer :: k
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
! note: do NOT add the extra names to profile_columns.list
! the profile_columns.list is only for the built-in profile column options.
! it must not include the new column names you are adding here.
! here is an example for adding a profile column
!if (n /= 1) stop 'data_for_extra_profile_columns'
!names(1) = 'beta'
!do k = 1, nz
! vals(k,1) = s% Pgas(k)/s% P(k)
!end do
end subroutine data_for_extra_profile_columns
integer function how_many_extra_history_header_items(id)
integer, intent(in) :: id
integer :: ierr
type (star_info), pointer :: s
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
how_many_extra_history_header_items = 0
end function how_many_extra_history_header_items
subroutine data_for_extra_history_header_items(id, n, names, vals, ierr)
integer, intent(in) :: id, n
character (len=maxlen_history_column_name) :: names(n)
real(dp) :: vals(n)
type(star_info), pointer :: s
integer, intent(out) :: ierr
ierr = 0
call star_ptr(id,s,ierr)
if(ierr/=0) return
! here is an example for adding an extra history header item
! also set how_many_extra_history_header_items
! names(1) = 'mixing_length_alpha'
! vals(1) = s% mixing_length_alpha
end subroutine data_for_extra_history_header_items
integer function how_many_extra_profile_header_items(id)
integer, intent(in) :: id
integer :: ierr
type (star_info), pointer :: s
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
how_many_extra_profile_header_items = 0
end function how_many_extra_profile_header_items
subroutine data_for_extra_profile_header_items(id, n, names, vals, ierr)
integer, intent(in) :: id, n
character (len=maxlen_profile_column_name) :: names(n)
real(dp) :: vals(n)
type(star_info), pointer :: s
integer, intent(out) :: ierr
ierr = 0
call star_ptr(id,s,ierr)
if(ierr/=0) return
! here is an example for adding an extra profile header item
! also set how_many_extra_profile_header_items
! names(1) = 'mixing_length_alpha'
! vals(1) = s% mixing_length_alpha
end subroutine data_for_extra_profile_header_items
! returns either keep_going or terminate.
! note: cannot request retry; extras_check_model can do that.
integer function extras_finish_step(id)
integer, intent(in) :: id
integer :: ierr
type (star_info), pointer :: s
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
extras_finish_step = keep_going
! to save a profile,
! s% need_to_save_profiles_now = .true.
! to update the star log,
! s% need_to_update_history_now = .true.
!! TASK block here:
!write(*,*) 'delta_Pg:', s% delta_Pg
if (s% delta_Pg > peak_delta_Pg) then
peak_delta_Pg = s% delta_Pg
write(*,*) 'New peak of delta_Pg found:', peak_delta_Pg
write(*,*) 'star age:', s%star_age
end if
!! TASK block ends here
! see extras_check_model for information about custom termination codes
! by default, indicate where (in the code) MESA terminated
if (extras_finish_step == terminate) s% termination_code = t_extras_finish_step
end function extras_finish_step
subroutine extras_after_evolve(id, ierr)
integer, intent(in) :: id
integer, intent(out) :: ierr
type (star_info), pointer :: s
ierr = 0
call star_ptr(id, s, ierr)
if (ierr /= 0) return
end subroutine extras_after_evolve
subroutine evaluate_cz_bdy_dq(s, cz_bdy_dq)
type(star_info), pointer, intent(in) :: s
real(dp), intent(out) :: cz_bdy_dq(s% nz)
real(dp) :: cz_dq
integer :: idx_bdy
cz_bdy_dq = 0d0
! We only do it for the first limit...
! Test and works!
idx_bdy = s%conv_bdy_loc(1)
cz_dq = find0(0._dp, s% gradr(idx_bdy+1) - s%gradL(idx_bdy+1),&
s%dq(idx_bdy), s% gradr(idx_bdy) - s% gradL(idx_bdy))
! If strange value (negative or bigger than dq) then don't interpolate and take the closest
! Probably not the best...
cz_bdy_dq(idx_bdy) = max(0d0, min(s% dq(idx_bdy) - cz_dq, s% dq(idx_bdy)))
end subroutine
! No diff with the overshoot_utils one
subroutine eval_conv_bdy_k_perso (s, i, k, ierr)
type(star_info), pointer :: s
integer, intent(in) :: i
integer, intent(out) :: k
integer, intent(out) :: ierr
! Evaluate the index k of the cell containing the i'th convective
! boundary
ierr = 0
if (s%top_conv_bdy(i)) then
k = s%conv_bdy_loc(i)
else
k = s%conv_bdy_loc(i) - 1
endif
if (k >= s%nz .OR. k < 1) then
write(*,*) 'Invalid cell for convective boundary: i, k, nz=', i, k, s%nz
ierr = -1
return
endif
! Finish
return
end subroutine eval_conv_bdy_k_perso
!****
subroutine eval_conv_bdy_r_perso (s, i, r, ierr)
type(star_info), pointer :: s
integer, intent(in) :: i
real(dp), intent(out) :: r
integer, intent(out) :: ierr
integer :: k
real(dp) :: w, cz_bdy_dq(s% nz)
! Evaluate the radius r at the i'th convective boundary
! Find the convective boundary cell
ierr = 0
call eval_conv_bdy_k_perso(s, i, k, ierr)
if (ierr /= 0) return
! Interpolate r based on the fact that r^3 varies linearly with q
! across the (constant-density) cell
call evaluate_cz_bdy_dq(s, cz_bdy_dq)
w = cz_bdy_dq(k)/s%dq(k)
if (w < 0._dp .OR. w > 1._dp) then
write(*,*) 'Invalid weight for convective boundary: i, w=', i, w
ierr = -1
return
end if
associate (k_o => k, &
k_i => k+1)
r = pow(( w)*s%r(k_i)*s%r(k_i)*s%r(k_i) + &
(1._dp-w)*s%r(k_o)*s%r(k_o)*s%r(k_o), 1._dp/3._dp)
end associate
! Finish
return
end subroutine eval_conv_bdy_r_perso
!****
subroutine eval_conv_bdy_Hp_perso (s, i, Hp, ierr)
type(star_info), pointer :: s
integer, intent(in) :: i
real(dp), intent(out) :: Hp
integer, intent(out) :: ierr
integer :: k
real(dp) :: r
real(dp) :: w
real(dp) :: x0
real(dp) :: x1
real(dp) :: x2
real(dp) :: x
real(dp) :: a0
real(dp) :: a1
real(dp) :: a2
real(dp) :: P
real(dp) :: rho
real(dp) :: r_top
real(dp) :: r_bot
real(dp) :: dr
real(dp) :: cz_bdy_dq(s% nz)
! Evaluate the pressure scale height Hp at the i'th convective boundary
! Find the convective boundary cell
ierr = 0
call evaluate_cz_bdy_dq(s, cz_bdy_dq)
call eval_conv_bdy_k_perso(s, i, k, ierr)
if (ierr /= 0) return
! Evaluate the radius at the convective boundary
call eval_conv_bdy_r_perso(s, i, r, ierr)
if (ierr /= 0) return
! Interpolate the pressure and density at the boundary, using a
! quadratic fit across the boundary cell and its neighbors (the
! x's are fractional mass distances from the outer edge of cell
! k-1); then, evaluate the pressure scale height
associate (k_o => k-1, &
k_m => k, &
k_i => k+1)
x0 = s%dq(k_o)/2._dp
x1 = s%dq(k_o) + s%dq(k_m)/2._dp
x2 = s%dq(k_o) + s%dq(k_m) + s%dq(k_i)/2._dp
x = s%dq(k_o) + cz_bdy_dq(k)
call two_piece_linear_coeffs(x, x0, x1, x2, a0, a1, a2, ierr)
if (ierr /= 0) return
P = exp(a0*s%lnPeos(k_o) + a1*s%lnPeos(k_m) + a2*s%lnPeos(k_i))
rho = exp(a0*s%lnd(k_o) + a1*s%lnd(k_m) + a2*s%lnd(k_i))
! Evaluate the pressure scale height
Hp = P/(rho*s%cgrav(k_m)* &
(s%M_center + s%xmstar*s%conv_bdy_q(i))/(r*r))
end associate
! (Possibly) limit the scale height using the size of the
! convection zone
if (s%limit_overshoot_Hp_using_size_of_convection_zone) then
! Determine the radial extent of the convection zone (note that
! r_top/r_bot don't coincide exactly with the r calculated
! above)
if (s%top_conv_bdy(i)) then
if (i == 1) then
r_bot = s%R_center
else
if (s%top_conv_bdy(i-1)) then
write(*,*) 'Double top boundary in overshoot; i=', i
ierr = -1
return
end if
r_bot = s%r(s%conv_bdy_loc(i-1))
endif
r_top = s%r(k)
else
r_bot = s%r(k+1)
if (i == s%num_conv_boundaries) then
r_top = s%r(1)
else
if (.NOT. s%top_conv_bdy(i+1)) then
write(*,*) 'Double bottom boundary in overshoot; i=', i
ierr = -1
return
endif
r_top = s%r(s%conv_bdy_loc(i+1))
endif
endif
dr = r_top - r_bot
! Apply the limit
if (s%overshoot_alpha > 0d0) then
if (s%overshoot_alpha*Hp > dr) Hp = dr/s%overshoot_alpha
else
if (s%alpha_mlt(k)*Hp > dr) Hp = dr/s%mixing_length_alpha
end if
end if
! Finish
return
end subroutine eval_conv_bdy_Hp_perso
!****
subroutine eval_over_bdy_params_perso (s, i, f0, k, r, D, vc, ierr)
type(star_info), pointer :: s
integer, intent(in) :: i
real(dp), intent(in) :: f0
integer, intent(out) :: k
real(dp), intent(out) :: r
real(dp), intent(out) :: D
real(dp), intent(out) :: vc
integer, intent(out) :: ierr
integer :: k_cb
real(dp) :: r_cb
real(dp) :: Hp_cb
real(dp) :: w
real(dp) :: lambda
! Evaluate parameters (cell index k, radius r, diffusion
! coefficients D and cdc) for the overshoot boundary associated
! with the i'th convective boundary
! Find the convective boundary cell
ierr = 0
call eval_conv_bdy_k_perso(s, i, k_cb, ierr)
if (ierr /= 0) return
! Evaluate the radius at the convective boundary
call eval_conv_bdy_r_perso(s, i, r_cb, ierr)
if (ierr /= 0) return
! Evaluate the pressure scale height at the convective boundary
call eval_conv_bdy_Hp_perso(s, i, Hp_cb, ierr)
if (ierr /= 0) return
! Search for the overshoot boundary cell
ierr = 0
if (s%top_conv_bdy(i)) then
! Overshooting outward -- search inward
r = r_cb - f0*Hp_cb
if (r <= s%r(s%nz)) then
r = s%r(s%nz)
k = s%nz - 1
else
search_in_loop: do k = k_cb, s%nz-1
if (s%r(k+1) <= r) exit search_in_loop
end do search_in_loop
endif
else
! Overshooting inward -- search outward
r = r_cb + f0*Hp_cb
if (r >= s%r(1)) then
r = s%r(1)
k = 1
else
search_out_loop : do k = k_cb, 1, -1
if (s%r(k) > r) exit search_out_loop
end do search_out_loop
endif
endif
if (.NOT. (s%r(k+1) <= r .AND. s%r(k) >= r)) then
write(*,*) 'r_ob not correctly bracketed: r(k+1), r, r(k)=', s%r(k+1), r, s%r(k)
ierr = -1
return
end if
! Interpolate mixing parameters
w = (s%r(k)*s%r(k)*s%r(k) - r*r*r)/ &
(s%r(k)*s%r(k)*s%r(k) - s%r(k+1)*s%r(k+1)*s%r(k+1))
lambda = (1._dp-w)*s%mlt_mixing_length(k) + w*s%mlt_mixing_length(k+1)
if (s%conv_vel(k) /= 0._dp .AND. s%conv_vel(k+1) /= 0._dp) then
! Both faces of cell have non-zero mixing; interpolate vc between faces
vc = (1._dp-w)*s%conv_vel(k) + w*s%conv_vel(k+1)
elseif (s%conv_vel(k) /= 0._dp .AND. s%conv_vel(k+1) == 0._dp) then
! Outer face of cell has non-zero mixing; interpolate vc
! between this face and r_cb, assuming vc = 0 at the latter
if(s%r(k) /= r_cb) then
w = (s%r(k)*s%r(k)*s%r(k) - r*r*r)/ &
(s%r(k)*s%r(k)*s%r(k) - r_cb*r_cb*r_cb)
else
w = 0d0
end if
vc = (1._dp-w)*s%conv_vel(k)
elseif (s%conv_vel(k) == 0._dp .AND. s%conv_vel(k+1) /= 0._dp) then
! Inner face of cell has non-zero mixing; interpolate vc
! between this face and r_cb, assuming vc = 0 at the latter
if(s%r(k+1) /= r_cb) then
w = (r_cb*r_cb*r_cb - r*r*r)/ &
(r_cb*r_cb*r_cb - s%r(k+1)*s%r(k+1)*s%r(k+1))
else
w = 0d0
end if
vc = w*s%conv_vel(k+1)
else
! Neither face of cell has non-zero mixing; return
vc = 0._dp
endif
! Evaluate the diffusion coefficient
D = vc*lambda/3._dp
! Finish
ierr = 0
return
end subroutine eval_over_bdy_params_perso
end module run_star_extrasnow, run your models:
./clean && ./mk
./rnA reminder from yesterday’s labs - if you want to save your terminal output to a text file, run:
./rn | tee out.txtThen, you can look for the last print of the peak_delta_Pg parameter in the terminal output and its corresponding stellar age. When you finish, don’t forget to fill in the values in the Google Sheet.
Note
As in Lab 2, you might see a warning message from MESA saying something like
WARNING: rel_run_E_err 13941 3.6538389571733347D+00Don’t worry, this is because the model is not very realistic (low resolution, simplified physics) in order to make it run faster.
The run should take around 10 minutes on 2 threads, since the star needs to evolve into the RC phase.
Additionally, we have prepared a Google Colab notebook. In this notebook, you can upload your MESA history.data file and generate plots of
versus age and central helium abundance. With these plots, you can see the evolution of
as a function of these parameters, not just their peak.
Instructions for the Google Colab notebook:
- Click here (local copy) to open the notebook and connect to your Google account.
- Make a copy of this notebook using
File -> Save a copy in Drive. - You can review the Python script if you’d like. You don’t need to install anything manually—just run the notebook cells, and it will automatically install any required packages. It uses the
mesa-readerpackage (more information here) to read the history file easily. - During the run (which will take 1–2 minutes), Colab will prompt you to upload a file. Please upload your
history.datafile when asked. - You will see the generated plot displayed below in the notebook.
Task 5: Create and Use a Custom Nuclear Reaction Network
We will learn how to create a new nuclear reaction network. We will then run the same inlists, but using the new reaction network that we create.
Up until now, we have used the basic.net network located in the $MESA_DIR/data/net_data/nets directory.
Copy the basic.net file to your working directory and give it a new name, like basic_custom.net, then open the file and review its contents.
The file includes two functions: add_isos() and add_reactions(). For more information about how these functions work and how to create a custom network, see the MESA documentation.
We will comment out all the helium-burning reactions except for: r_he4_he4_he4_to_c12 and r_c12_ag_o16.
This is what your basic.net file should look like
! the basic net is for "no frills" hydrogen and helium burning.
! assumes T is low enough so can ignore advanced burning and hot cno issues.
add_isos(
h1
he3
he4
c12
n14
o16
ne20
mg24
)
add_reactions(
! pp chains
rpp_to_he3 ! p(p e+nu)h2(p g)he3
rpep_to_he3 ! p(e-p nu)h2(p g)he3
r_he3_he3_to_h1_h1_he4 ! he3(he3 2p)he4
r34_pp2 ! he4(he3 g)be7(e- nu)li7(p a)he4
r34_pp3 ! he4(he3 g)be7(p g)b8(e+ nu)be8( a)he4
r_h1_he3_wk_he4 ! he3(p e+nu)he4
! cno cycles
rc12_to_n14 ! c12(p g)n13(e+nu)c13(p g)n14
rn14_to_c12 ! n14(p g)o15(e+nu)n15(p a)c12
rn14_to_o16 ! n14(p g)o15(e+nu)n15(p g)o16
ro16_to_n14 ! o16(p g)f17(e+nu)o17(p a)n14
! helium burning
r_he4_he4_he4_to_c12
r_c12_ag_o16
!rc12ap_to_o16 ! c12(a p)n15(p g)o16
!rn14ag_lite ! n14 + 1.5 alpha = ne20
!r_o16_ag_ne20
!ro16ap_to_ne20 ! o16(a p)f19(p g)ne20
!r_ne20_ag_mg24
!rne20ap_to_mg24 ! ne20(a p)na23(p g)mg24
! auxiliaries -- used only for computing rates of other reactions
rbe7ec_li7_aux
rbe7pg_b8_aux
rn14pg_aux
rn15pg_aux
rn15pa_aux
ro16ap_aux
rf19pg_aux
rf19pa_aux
rne20ap_aux
rna23pg_aux
rna23pa_aux
rc12ap_aux ! c12(ap)n15
rn15pg_aux ! n15(pg)o16
ro16gp_aux ! o16(gp)n15
rn15pa_aux ! n15(pa)c12
)Edit your inlist_project to include the new network - you can find the relevant inlist parameters in the MESA documentation we referred you to before.
Solution 5
! new net
change_net = .true.
change_initial_net = .true.
new_net_name = 'basic_custom.net'MESA will look for basic_custom.net both in your working directory and in $MESA_DIR/data/net_data/nets.
Run and re-plot
versus age and central helium abundance using the Google Colab notebook again (you will need to upload your new history.data file). You can open a new window of the Google Colab Click here
(local copy)
, or save the previous plot for comparison.
Not much has changed, right?
This shows us that the helium-burning reaction processes that most influence the period spacing—and therefore the interior structure of the star—are: r_he4_he4_he4_to_c12 and r_c12_ag_o16.
If you comment one of them out (just for the exercise—it doesn’t come from a real physical motivation), you will already start to see a more noticeable change in the period spacing plots, and hence, also in the interior structure of the star.
This task took inspiration from one of last year’s MESA Summer School labs. If you want to learn more about changing nuclear networks in MESA, please look here.
Task 6 - Bonus: Add Plots for Abundances and Temperature-Density Evolution
We can add two plots that will give us information about the reaction rates and the temperature changes during the run.
Replace the Kippenhahn diagram and the mixing profile plot you added in Maxilab1 to make more space for our two new plots, with the following:
- An abundance plot of all the elements in the nuclear reaction network we are using, displayed as mass fraction (in log scale) as a function of the mass profile of the star.
- The central temperature ( ) as a function of the central density ( ) in log-log scale.
Hint 6
These plots are included in the MESA pgstar defaults and can be added easily.
- Look in the MESA documentation under Using MESA / Using PGSTAR, where you will find The Inventory of Plots section with the available plot titles.
- You can also check the
$MESA_DIR/star/defaults/pgstar.defaultsfile to see the default plot settings and titles you can use.
Solution 6
- Replace this line:
Grid1_plot_name(1) = 'Kipp'with:Grid1_plot_name(1) = 'Abundance'. - Replace this line:
Grid1_plot_name(5) = 'Mixing'with:Grid1_plot_name(5) = 'TRho'.
Your complete inlist_pgstar should look like this:
&pgstar
! Set up grid layout
file_white_on_black_flag = .false.
Grid1_win_flag = .true.
Grid1_file_interval = 5
Grid1_file_width = 25
Grid1_num_cols = 10
Grid1_num_rows = 10
Grid1_win_width = 14
Grid1_win_aspect_ratio = 0.5
Grid1_xleft = 0.00
Grid1_xright = 1.00
Grid1_ybot = 0.00
Grid1_ytop = 1.00
Grid1_num_plots = 8
! Add Abundance plot
Grid1_plot_name(1) = 'Abundance'
Grid1_plot_row(1) = 1
Grid1_plot_rowspan(1) = 5
Grid1_plot_col(1) = 1
Grid1_plot_colspan(1) = 4
Grid1_plot_pad_left(1) = 0.05
Grid1_plot_pad_right(1) = 0.01
Grid1_plot_pad_top(1) = 0.04
Grid1_plot_pad_bot(1) = 0.05
Grid1_txt_scale_factor(1) = 0.5
! Add HR diagram
Grid1_plot_name(2) = 'HR'
Grid1_plot_row(2) = 6
Grid1_plot_rowspan(2) = 3
Grid1_plot_col(2) = 1
Grid1_plot_colspan(2) = 2
Grid1_plot_pad_left(2) = 0.05
Grid1_plot_pad_right(2) = 0.01
Grid1_plot_pad_top(2) = 0.02
Grid1_plot_pad_bot(2) = 0.07
Grid1_txt_scale_factor(2) = 0.5
! Add abundance profile plot
Grid1_plot_name(3) = 'Power'
Grid1_plot_row(3) = 6
Grid1_plot_rowspan(3) = 3
Grid1_plot_col(3) = 3
Grid1_plot_colspan(3) = 2
Grid1_plot_pad_left(3) = 0.05
Grid1_plot_pad_right(3) = 0.01
Grid1_plot_pad_top(3) = 0.02
Grid1_plot_pad_bot(3) = 0.07
Grid1_txt_scale_factor(3) = 0.5
! Add text panel
Grid1_plot_name(4) = 'Text_Summary1'
Grid1_plot_row(4) = 9
Grid1_plot_rowspan(4) = 2
Grid1_plot_col(4) = 1
Grid1_plot_colspan(4) = 10
Grid1_plot_pad_left(4) = 0.00
Grid1_plot_pad_right(4) = 0.00
Grid1_plot_pad_top(4) = 0.00
Grid1_plot_pad_bot(4) = 0.00
Grid1_txt_scale_factor(4) = 0
Text_Summary1_name(1,1) = 'model_number'
Text_Summary1_name(2,1) = 'star_age'
Text_Summary1_name(3,1) = 'log_dt'
Text_Summary1_name(4,1) = 'luminosity'
Text_Summary1_name(5,1) = 'Teff'
Text_Summary1_name(6,1) = 'radius'
Text_Summary1_name(7,1) = 'log_g'
Text_Summary1_name(8,1) = 'star_mass'
Text_Summary1_name(1,2) = 'log_cntr_T'
Text_Summary1_name(2,2) = 'log_cntr_Rho'
Text_Summary1_name(3,2) = 'log_center_P'
Text_Summary1_name(4,2) = 'center h1'
Text_Summary1_name(5,2) = 'center he4'
Text_Summary1_name(6,2) = ''
Text_Summary1_name(7,2) = ''
Text_Summary1_name(8,2) = ''
Text_Summary1_name(1,3) = 'log_Lnuc'
Text_Summary1_name(2,3) = 'log_Lneu'
Text_Summary1_name(3,3) = 'log_LH'
Text_Summary1_name(4,3) = 'log_LHe'
Text_Summary1_name(5,3) = 'num_zones'
Text_Summary1_name(6,3) = ''
Text_Summary1_name(7,3) = ''
Text_Summary1_name(8,3) = ''
Text_Summary1_name(1,4) = 'nu_max'
Text_Summary1_name(2,4) = 'delta_nu'
Text_Summary1_name(3,4) = 'delta_Pg'
Text_Summary1_name(4,4) = ''
Text_Summary1_name(5,4) = ''
Text_Summary1_name(6,4) = ''
Text_Summary1_name(7,4) = ''
Text_Summary1_name(8,4) = ''
! Add temperature-density plot
Grid1_plot_name(5) = 'TRho'
Grid1_plot_row(5) = 1
Grid1_plot_rowspan(5) = 4
Grid1_plot_col(5) = 6
Grid1_plot_colspan(5) = 4
Grid1_plot_pad_left(5) = 0.05
Grid1_plot_pad_right(5) = 0.05
Grid1_plot_pad_top(5) = 0.04
Grid1_plot_pad_bot(5) = 0.07
Grid1_txt_scale_factor(5) = 0.5
Grid1_file_flag = .true.
Grid1_file_dir = 'pgplot'
Grid1_file_prefix = 'grid_'
file_white_on_black_flag = .true.
Grid1_file_interval = 10
! Add mode inertia panel
Grid1_plot_name(6) = 'History_Panels1' !
Grid1_plot_row(6) = 5
Grid1_plot_rowspan(6) = 4
Grid1_plot_col(6) = 6
Grid1_plot_colspan(6) = 5
History_Panels1_win_flag = .true.
History_Panels1_num_panels = 2
History_Panels1_xaxis_name = 'model_number'
History_Panels1_yaxis_name(1) ='delta_Pg'
History_Panels1_other_yaxis_name(1) = ''
History_Panels1_yaxis_name(2) ='center_he4'
History_Panels1_other_yaxis_name(2) = ''
History_Panels1_automatic_star_age_units = .true.
Grid1_plot_pad_left(6) = 0.05
Grid1_plot_pad_right(6) = 0.05
Grid1_plot_pad_top(6) = 0.04
Grid1_plot_pad_bot(6) = 0.07
Grid1_txt_scale_factor(6) = 0.5
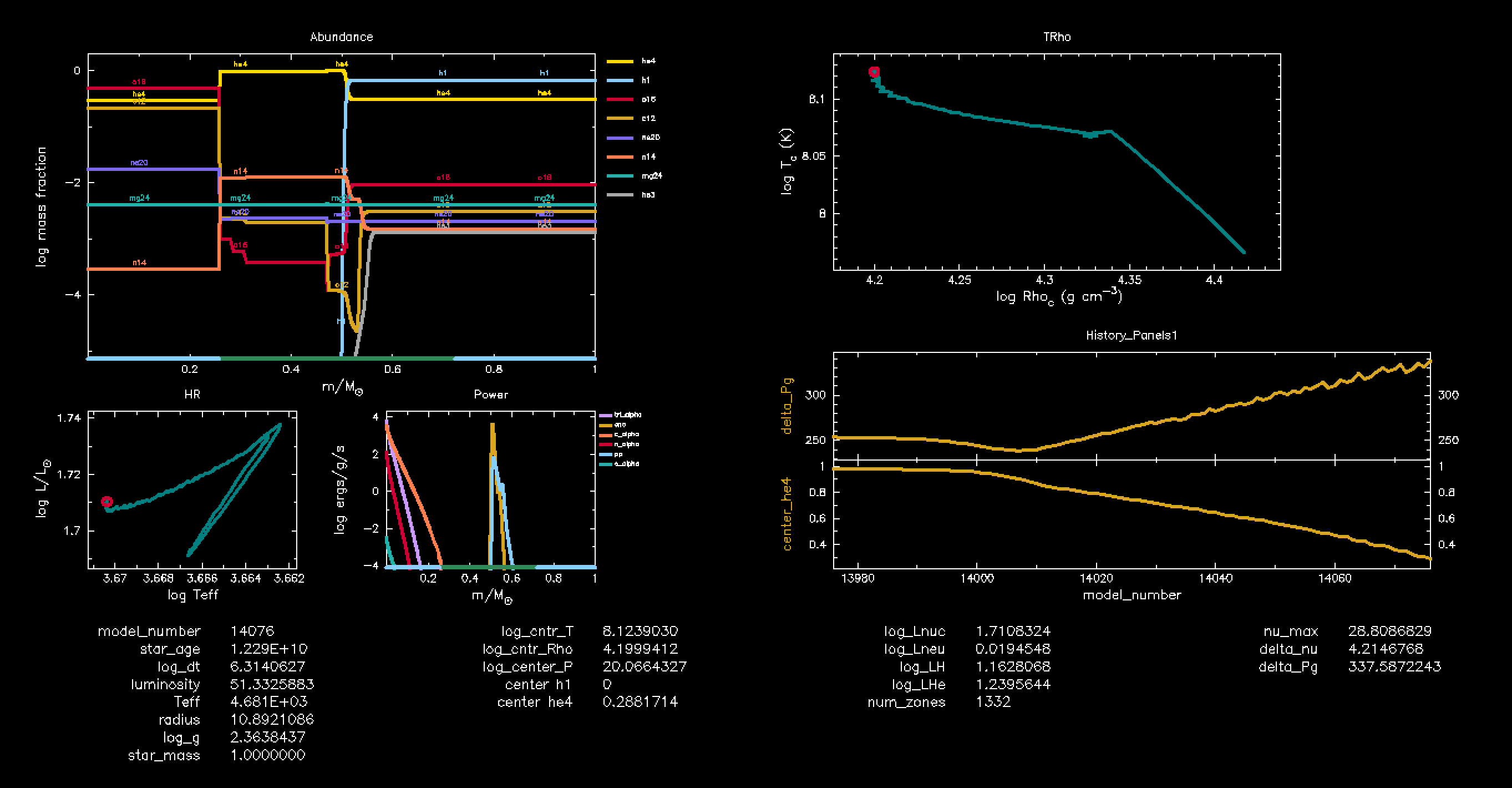
/ ! end of pgstar namelistNow run your models. In the pgplot window, you will be able to observe the changing abundances of the elements involved in He-burning (mainly He4, Be7, C12, and O16) as the temperature of the star changes.
This is how your pgplot window is supposed to look: