Monday MaxiLab 1: Modeling core overshooting in main-sequence stars
In this lab, you will learn how to set up a MESA model from scratch, monitor the run, customize its output and choose reasonable values for model parameters. Our science case is focused on the effects of core overshooting on the core of main-sequence stars.
As a maxilab, this lab will take two 1.5h sessions. Each session is organized in three sections, structured as below with an estimate of the time you are expected to spend on each. Please do not hesitate to ask your TA and/or the other people at your table for assistance if you notice you are falling behind.
Session 1
Setting up your MESA work directory [~20 min.]
Copying the default work dir
We will start from the default MESA work directory and slowly build it up until we have a fleshed-out main-sequence model. Copy the default work directory into a new folder called lab1 and navigate into said folder:
cp -r $MESA_DIR/star/work lab1
cd lab1Do a quick ls to check what is included in this default work directory.
You’ll see a number of executables, namely clean, mk, re and
rn. The subdirectories make and src contain the
Makefile and extra code to include, but you don’t have to look into that
today. For now, let’s take a look at the inlists inlist,
inlist_pgstar and inlist_project.
The term inlist refers to MESA’s standard Fortran namelist file that launches a simulation.
It’s a plain text file where you group configuration variables into sections (namelists)
These files describe what you want MESA to do. In particular MESA will always look for inlist.
Using your favorite text editor, take a look at what is in inlist.
What this inlist essentially does is redirect MESA to the other two inlist files for all the real content, with inlist_project containing most of the fields describing how the MESA run should go and inlist_pgstar describing what visuals MESA should produce. For now, focus on inlist_project.
Basic initial parameters
Let’s start with a very simple main-sequence model of a star with an initial mass of 5 solar masses and metallicity of 0.014 with some strong step-wise mixing due to core overshooting. To do so, open inlist_project and find and change the following parameters to the given values:
initial_mass = 5d0
initial_z = 0.014d0Note
Have you spotted those d0 at the end of these lines? The d therein indicates that the numbers we provide are double precision floating point numbers in fortran. The number afterwards, 0 in this case, indicates the order of magnitude in a scientific notation. For example, 2.2d3 = 2200.0 or we could have written initial_z = 1.4d-2. Even when the order is zero, it is good practice to always add d0 after your floats.
The opacity table MESA uses depends on the reference metallicity,
defined by Zbase under the &kap namelist.
For consistency, you should set Zbase to the same value as
initial_z:
Zbase = 0.014d0Adding overshooting
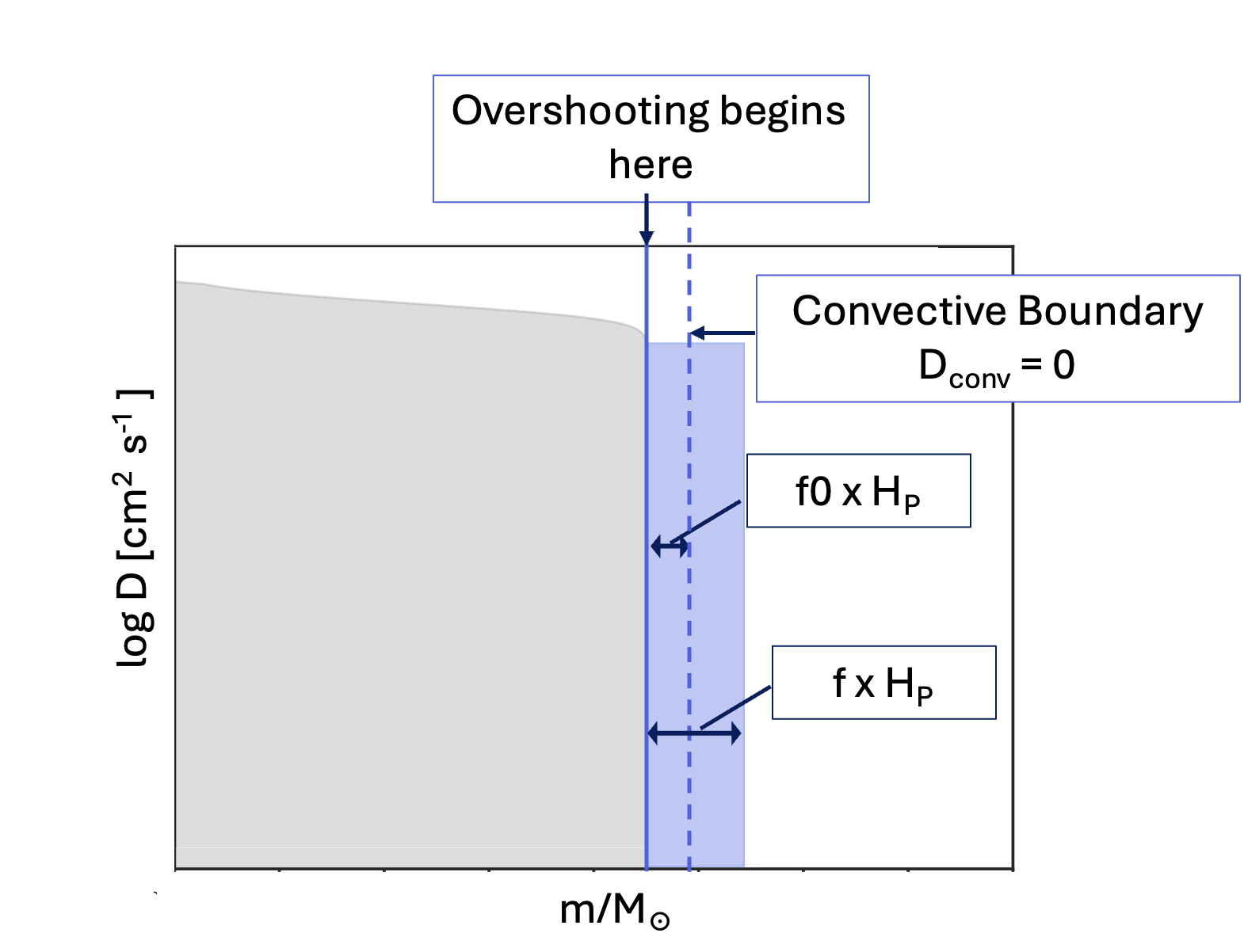
Next, to add the core overshooting, we need to add in some new fields, shown below. As you plug this into your inlist, have a look at the questions below.
overshoot_zone_type(1) = 'burn_H'
overshoot_zone_loc(1) = 'core'
overshoot_bdy_loc(1) = 'top'
overshoot_scheme(1) = 'step'
overshoot_f(1) = 0.30d0
overshoot_f0(1) = 0.005d0Question: The first three overshoot_ fields describe where the overshooting should take place. Go into the MESA documentation and look up what each of these fields means.
Click here to show the answer
overshoot_zone_typelets you only activate overshooting around regions where certain types of burning takes place.overshoot_zone_locthen tells MESA whether this overshoot zone should surround the convective core or a shellovershoot_bdy_locwhether the overshoot should occur only above or only below the relevant convection zone(s).
Question: Where should you add these fields?
Click here to show the answer
&controls namelist. However, you’ll probably notice that &controls is organized into subsections like “starting specifications”, “when to stop”, “wind” and so on. Sticking to this or a similar structure is a good idea to keep your inlist clearly organized.
As such, we recommend copy-pasting the six lines above under “! mixing”.Bonus Question: Why does each overshoot field in our example have that (1) at the end?
Click here to show the answer
overshoot_ are actually arrays and (1) indicates the first element of that array in fortran. This way, each element can represent a different overshooting zone so you can use different settings for each overshooting zone.Termination conditions
Before you run your model, you should consider when the model is
terminated. Since we want to simulate the main-sequence evolution,
we should place our stopping condition at the terminal age
main-sequence (TAMS). Look under ! when to stop in &controls
of your inlist_project.
There are two conditions that can trigger the model to end. The first is
Lnuc_div_L_zams_limit = 0.99d0
stop_near_zams = .true.which is designed to stop the model at the zero-age main-sequence (ZAMS),
that it defines as the point where 99% of the energy released comes from
nuclear reactions. As we seek to model the main-sequence, we obviously
do not want the run to end around the ZAMS. Therefore, disable stop_near_zams by setting:
stop_near_zams = .false.The second condition is meant to stop the model around the TAMS.
xa_central_lower_limit_species(1) = 'h1'
xa_central_lower_limit(1) = 1d-3Question: How does the default inlist_project define the TAMS?
Click here to show the answer
For our purposes today, it will be interesting to go a little bit further. Therefore, change the lower limit on H to :
xa_central_lower_limit_species(1) = 'h1'
xa_central_lower_limit(1) = 1d-6Running the model
Run your model to the TAMS by cleaning any executables in your work directory using
./clean
./mk
./rnAs your model runs, you will notice that MESA writes heaps of numbers to your terminal. After a while, two panels with constantly changing plots pop up. They show the evolutionary track of your model on a Hertzsprung-Russell diagram and one the internal temperature and density profiles. These help you keep a close eye on your model and can help you identify problems and potential improvements.
For now, let’s focus on the terminal output. One important field for our
purposes is the central hydrogen fraction (H_cntr), which tells
you how far along the main-sequence evolution your model is. Note
that it initially changes extremely slowly. This is because MESA starts
with a very small time step which gradually increases, as shown by the
lg_dt_yrs field, the base 10 logarithm of the time step expressed in
years.
Some optimisations
Before running our main model, let’s make two adjustments to speed up the MESA run:
MESA starts by creating a pre-main-sequence model (before nuclear burning begins) and by default performs 300 relaxation steps to ensure it’s stable. For our educational purposes, we can reduce this to save time by adding to the &star_job section of your inlist_project:
pre_ms_relax_num_steps = 100MESA also starts with very tiny time steps after the pre-main sequence model has been created and relaxed. We can tell it to use larger initial steps, by adding the following under &star_job:
set_initial_dt = .true.
years_for_initial_dt = 1d0These adjustments will help our model run faster, especially during the early phases. In your future research, you’d want to carefully test how these parameters affect your specific science case, but for this lab, these settings are fine.
If you want to (and are on schedule), you can briefly run your model
again by entering ./rn in your terminal to test whether
your changes to the inlist did what they are supposed to.
Tip
You don’t have to run your model all the way to the TAMS. You can interrupt it using ctrl+C.
If you got stuck and cannot get your inlist to run, you can find a functional inlist with all the changes described above here , so you can continue with limited delay. Be sure to rename inlist_project1 to inlist_project!
Modifying the input physics and saving your final model [~20 min.]
Mass loss
inlist_project is currently mostly empty, meaning most settings are using MESA’s default values. You should always check whether these are appropriate for your models. As an example, massive stars can loose a fair amount of mass through winds.
Check what the default wind mass loss is.
You can find this in the documentation
of &controls to see what the default wind mass loss is. You could also look through
the default values in the MESA code inside $MESA_DIR/star/defaults/ (controls.defaults in this case), though the website
documentation is easier to navigate if you don’t know the name of the relevant fields yet.
Click here to show a hint
In the panel on the left of the website, navigate to Reference and Defaults > controls. On the right, you can now see the contents of this page. Mass loss by winds is found under mass gain and loss.
Tip
All apart from the most recent MESA build, this is the path to controls: “Reference > star defaults > controls” If you are using the most recent MESA build then the path is “Reference and Defaults > controls”
Click here to show the answer
mass_change or with some wind_scheme.You will see in the documentation that there is a wealth of wind mass loss schemes available, all of which can be scaled up or down by a constant factor. Each scheme is appropriate in particular regimes of the surface temperature, composition, etc. The so-called Dutch scheme attempts to merge some of these schemes into a cohesive whole. Add it into your inlist_project without scaling.
Click here to show how to implement this
In order to use the Dutch scheme at all temperature ranges and without changing its scaling, use:
hot_wind_scheme = 'Dutch'
cool_wind_RGB_scheme = 'Dutch'
Dutch_scaling_factor = 1d0The mixing-length parameter
MESA uses the mixing-length theory (MLT) to describe the transport by convection. This theory relies on the scaling factor , which expresses for how many pressure scale heights a convective fluid parcel maintains its properties. Typically values range between 1 and 3. You should check what MESA’s default value of this parameter is.
Tip
The field setting
is called mixing_length_alpha. You can easily navigate to its description by entering this field name into the search bar on the top left of the MESA documentation website. If you do so, it’s best to click on controls > mixing_length_alpha since that will take you straight to the description of mixing_length_alpha.
When you are working on your real science cases, you should test a few different values for this to gain an understanding of its effects. However, to save some time in this lab, we will stick to just one value, namely 1.8. Add this into your inlist_project under &controls:
mixing_length_alpha = 1.8d0Saving the final model
In the other labs today, you will learn how to run models that continue after the main-sequence evolution. Instead of re-running the main-sequence evolution every time you tweak a setting, we can tell MESA to save a model at the end of a main-sequence run.
The default inlist explicitly tells MESA not to save the model. Find the fields below and change them to:
save_model_when_terminate = .true.
! Give a name to the model file to be saved including your parameter values, e.g.
! 'M{your_M}_Z{your_Z}_fov{your_f_overshoot}_f0ov{your_f0_overshoot}.mod'
save_model_filename = 'M5_Z0014_fov030_f0ov0005_TAMS.mod'Running the updated model
Despite how much you already added into your inlist_project, there are still many empty headers. Indeed, when building an inlist for your real science cases, you should still look into your opacity tables, atmosphere settings, equation of state, spatial and temporal resolution, and much more. However, for the sake of time, we’ll stop here.
Now let’s run the model again! To do so, enter
./rninto your terminal.
Important
When the run is finished, double check if the new file ‘M5_Z0014_fov030_f0ov0005_TAMS.mod’ is in your work directory. You will need that model for the next lab! If it’s not there, ask your TA for help.
If you got stuck and cannot get your inlist to run, you can find a functional inlist with all the changes described above here , so you can continue with limited delay. Be sure to rename inlist_project2 to inlist_project!
Customising output [~30 min.]
Upgrading the pgstar plots
Now let’s turn to these animated plots, often called the pgstar plots. These are incredibly useful in understanding what is going on in your model while its running, helping you spot potential problems early. Therefore, it is worthwhile to customize your pgstar panels to show those quantities that are the most important to your work. To this end, MESA has a bunch of prepared windows you can easily add by adding one flag to your inlist_pgstar. You can find these and how to edit your inlist_pgstar in this documentation page.
For this lab, we have prepared a specialized
inlist_pgstar for you. **Download that inlist_pgstar **
here
and move it into your MESA work directory. Make sure to name the file inlist_pgstar!
Run your model again to see what the new pgstar plots look like.
Tip
You don’t have to wait for the run to be finished. Remember that you can interrupt it using ctrl+C.
For some of you, this new panel may be very small or bigger than your screen. This is because the width of the pgstar window is dependent on your system and the size of your screen. If the panel is too large or small for you, open inlist_pgstar, find the two lines shown below near the top of the inlist and edit these values until the panel looks nice.
Grid1_win_width = 10
Grid1_win_aspect_ratio = 0.7Tip
You can edit inlist_pgstar while the model is running and it will immediately update your plots.
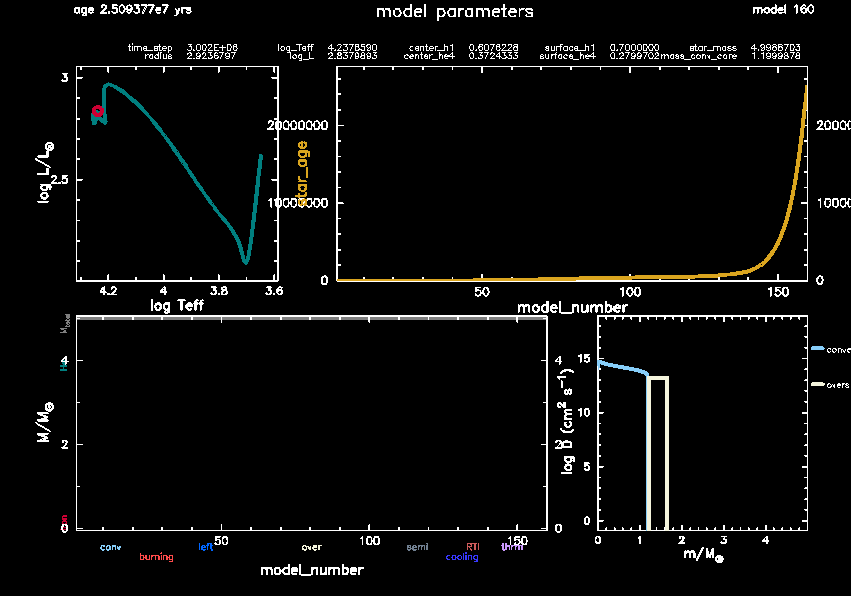
Understanding this pgstar window
We have merged all the plots in one panel for a better overview. We also included some key quantities at the top, similar to MESA’s terminal output.
The plots are:
- Top left: The Hertzsprung-Russell diagram (HRD),
- Top right: History panel showing model number on the x-axis and age on the y-axis.
(This is often used as a proxy for age since model number increments regularly during evolution), - Bottom right: The mixing panel,
- Bottom left: This is a mysterious, mostly-empty panel. We’ll get back to that empty panel later.
We’ll refer to the “history plot” (top right) throughout the lab, so keep that panel in mind.
Tip
inlist_pgstar saves the pgstar plots in the png directory, so you can look in there after MESA closed the pgstar panel at the end of the run. You could also tell your inlist_project not to close the pgstar panel until you tell it to. To do so, add the following into your inlist_project under star_job
pause_before_terminate = .true.For now, turn to that mixing panel in the bottom right and ponder these questions:
Question: What is the mixing panel showing exactly? What does the color of each line indicate?
Click here to show the answer
Bonus Question: Why do we plot the model number on the x-axis instead of, say, the stellar age?
Click here to show the answer
Customizing MESA’s output data
In your pgstar window, look at the plot in the top right corner. This panel is showing how model properties change over time, currently using model number as the x-axis (In many cases, the absolute age of the star doesn’t matter. Since age correlates closely with model number, and model number is easier to interpret, we often use it as a proxy). Since we’re studying core overshooting effects, we’d like to modify this panel to show the evolution of the core mass instead of the default value.
To customize what’s displayed in MESA plots, we first need to understand what data is being tracked. When MESA runs, it creates a LOGS directory containing two main types of output files:
history.data: Records global stellar properties at each timestepprofile*.data: Captures the star’s internal structure at specific timesteps
Open LOGS/history.data with a text editor. After the header information, you’ll see a list of column names around line 6.
These are all the quantities MESA has been tracking during your run.
To add more quantities to track, we need to customize the history columns list. First, we need to copy the default history column list to our work folder:
cp $MESA_DIR/star/defaults/history_columns.list my_history_columns.listOpen my_history_columns.list and take a minute to explore the wealth of available output options.
The file is organized by category, with comments explaining most variables.
This gives you a comprehensive view of what MESA can track.
For our overshooting study, we’re interested in the stellar core properties.
The convective core mass is already included in the defaults.
Let’s add the star’s radius to the history output by finding and uncommenting the radius field in the appropriate section.
Once you have uncommented the relevant lines, you need to tell MESA that you want to include the output columns in my_history_columns.list. To do so, add the following line into your inlist_project under &star_job:
history_columns_file = 'my_history_columns.list'Finally, let’s consider how often MESA writes this history output.
Bonus Question: How often does MESA write its history output by default?
Click here to show the answer
history_interval in the documentation, which is 5 by default.As we don’t need that many history columns, it’s reasonable to write them at
every step. Do so by setting in your &controls:
history_interval = 1Configuring the History Plot
Now let’s customize the History Panel in the pgstar window. This panel shows how stellar properties evolve over time.
- Open
inlist_pgstarin your text editor - Find the section that defines the History Panel (search for “History_Panels1”)
- Change the y-axis to display the convective core mass:
History_Panels1_yaxis_name(1) = 'mass_conv_core'This will show how the convective core mass evolves throughout the star’s main sequence lifetime.
To enable more advanced plotting features like the Kippenhahn diagram (which shows the internal structure evolution), we need to tell MESA to track additional data. Open my_history_columns.list and find and uncomment the following options:
mixing_regions 20
burning_regions 20These settings tell MESA to track up to 20 distinct mixing and nuclear burning regions within the star, which enables the creation of Kippenhahn diagrams and other detailed evolutionary plots.
Run your model briefly to confirm your pgstar window now displays the updated plot configuration.
Bonus Task – Profile Data
While history files track global properties over time, profile files capture the star’s internal structure at specific moments. These are essential for examining how variables change with radius inside the star. Copy over the default profile column list.
cp $MESA_DIR/star/defaults/profile_columns.list my_profile_columns.listTo study mixing processes in the stellar interior, uncomment these fields:
log_D_mix- the diffusion coefficient for mixingmixing_type- identifies which mixing process is active in each zone
After modifying this file, tell MESA to use your custom profile columns by adding to your inlist_project under &star_job:
profile_columns_file = 'my_profile_columns.list'Bonus Question: By default, MESA only saves a profile every 50 model steps. This might miss important evolutionary phases. How could you increase this frequency?
Click here to show the answer
profile_interval = 10 to your inlist’s &controls section to save profiles more frequently. You could also enable write_profile_when_terminate = .true. to guarantee a profile at the end of the run.Bonus Question: How many profiles can MESA write over the course of a run? Why might you want to limit it?
Click here to show the answer
max_num_profile_models in &controls). If the run keeps going after the maximum has been reached, the oldest profiles are overwritten.If you got stuck and cannot get your inlist to run, you can find a functional inlist with all the changes described above here , so you can continue with limited delay. Be sure to rename inlist_project3 to inlist_project! You can also find the completed inlist pgstar here, the completed history columns list here, and the completed profile columns list here.
Bonus Task – Movies
Bonus Task – Movies
Making an animation with the png/ folder:
Students who finish early can create an animation with the images in the png folder:
- Ensure
Grid1_file_flag = .true.is set (it already is in the completed inlist) - Run a quick model to generate PNGs
- Verify the outputs
ls -lh png/You should see quite a few png images.
- Now we can create the video: Move to the directory
cd pngMESA includes a script that is able to create movies. More instructions.
images_to_movie '*.png' evolution.mp4If this fails to find the .png files, try replacing the single quotation marks with double quotation marks, i.e.
images_to_movie "*.png" evolution.mp4Warning
If you’re on a Mac, this script to make movies may not work, depending on which MESA SDK you have installed. If you happen to have an older version of the MESA SDK, you can try initialising that and running the movies again. Otherwise, this bonus task will probably not work for you, so feel free to skip it or follow along with someone else.
Session 2
Adapting the input parameters and assessing their importance [~15 min.]
You now know how to navigate your work directory and build up a main-sequence model. That’s great! However, so far we have limited ourselves to simply adding in pre-chosen parameter values, choices of tables etc. In real scientific applications, you should always consider the impact of these settings, for instance by trying a few different values. In particular, there are a number of numerical schemes and uncertain physical parameters for which you should think carefully about the appropriate value. You already encountered some of these today, namely the mixing length parameter and convective overshooting.
In this session, we’ll explore the impact of overshooting in your model. The plan is that everyone gets a unique set of overshooting parameters, initial mass and initial metallicity to try out. You will then compare the results of these parameter settings to the model you produced in session 1. Meanwhile, we will collect some basic results from everyone’s models and examine the correlations between different parameters together.
Bonus Question: Overshooting in MESA starts overshoot_f0 scale heights into the convection zone. If you use step overshooting, the overshooting ends overshoot_f scale heights above the start of overshooting, after which the overshoot mixing goes to zero. But how does MESA define where the overshooting stops when you use exponential overshooting, which never goes all the way to zero?
Click here to show the answer
Picking new initial parameters
Go into this spreadsheet (mirror) and put your name next to one set of parameters to claim it as yours. Modify your inlist accordingly.
Since there are more parameter sets available than there are students, you can test different parameters if time allows. However, the main focus should be on completing this task and fully understanding it. Additionally, the bonus tasks in this lab involve running this set of parameters in a batch.
If you selected the ’no overshoot’ scheme from the spreadsheet, you should leave the overshoot scheme as an empty string, i.e.
overshoot_scheme(1) = ''If you selected the ’exponential’ scheme, you should consider where the overshoot region ends. As the mixing strength decreases exponentially, it approaches zero far away from the core-envelope boundary. As such, past a certain point, the mixing is so small as to become meaningless. Therefore, MESA stops the overshoot mixing when the mixing coefficient drops below 100 cm
/s. This is quite a high lower limit, so reduce it by setting in your &controls :
overshoot_D_min = 1d-2Preserving Your Previous Results
Caution
Before running a new model with different parameters, we need to ensure we don’t overwrite our previous results.
We’ll make two adjustments to keep our work organized. After selecting your parameter set from the spreadsheet, modify the save_model_filename in your inlist_project file to reflect these specific parameters:
save_model_filename = 'M{mass}_Z{metallicity}_{scheme}_fov{fov}_f0ov{f0}.mod'For example, if you selected a 15 star with , exponential overshooting with and , your filename would be:
save_model_filename = 'M15_Z0014_exponential_fov001_f0ov0001.mod'MESA normally writes all history and profile data to a directory called LOGS/. To keep these outputs separate from your previous run, we’ll direct MESA to use a different output directory. Add the following to your inlist_project under the &controls section:
log_directory = 'LOGS_M{mass}_Z{metallicity}_{scheme}_fov{fov}_f0ov{f0}'Using the same example parameters as above:
log_directory = 'LOGS_M15_Z0014_exponential_fov001_f0ov0001'This keeps all your runs organized with descriptive names that make it easy to identify which parameters were used for each model.
Running your model again
Now run your model again. Keep a close eye on your pgstar plots, particularly the mixing panel. Compare it with those of the other people at your table.
Collecting the TAMS output
After your model finishes running, we’ll extract key parameters at the Terminal Age Main Sequence (TAMS).
Open the final history.data file in your new LOGS directory with a text editor. The TAMS is represented by the final line in this file - this is where the central hydrogen has been depleted to the threshold we set (1d-6) and the run was ended.
From this final line, record the following values in the second page of the spreadsheet:
- log_Teff (logarithm of effective temperature)
- log_L (logarithm of luminosity in solar units)
- mass_conv_core or he_core_mass (depending on which is available)
- conv_mx1_top_r (if you uncommented this)
- star_age/1e6 (to convert to Myr)
These values will allow us to analyze how different overshooting parameters affect stellar evolution.
If you got stuck and cannot get your inlist to run, you can find a functional inlist with all the changes described above here , so you can continue with limited delay. Be sure to rename inlist_project4 to inlist_project! You will still have to update the parameters and output filenames and directory according to your parameter set though. Ask your TA for assistance with that.
Bonus Task – Plotting MESA output
Now let’s wrap up this lab by reading your MESA output in using Python and making some custom plots.
For an introduction into utilising python as an analysis tool, see bonus_tasks/python_analysis.
Bonus Task – Batch Parameter Studies with MESA
If you’ve completed the main lab activities and have time remaining, explore the automated parameter study framework in the bonus_tasks/ directory. This framework enables systematic exploration of overshooting effects across a grid of stellar model parameters.
For complete documentation and additional analysis tools, see bonus_tasks/.